Components
Metapsy consists of a collection of software tools and websites. Together, this infrastructure provides an integrated meta-analysis workflow.
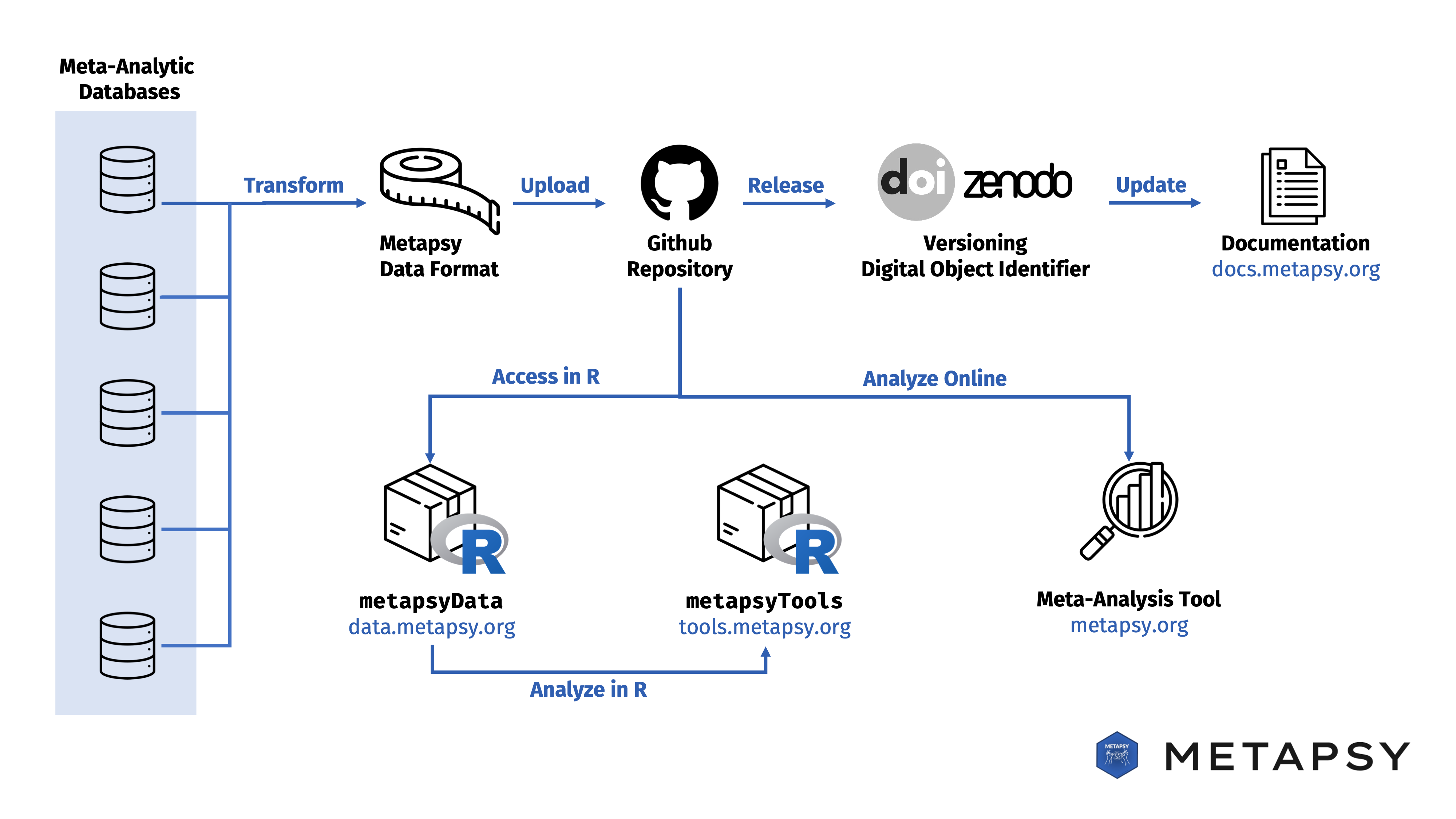
Components of the Metapsy data flow:
- Meta-analytic databases are maintained and regularly updated by specialized university-based research teams.
- Updated or newly created databases are then transformed into a consistent format (Metapsy data format).
- Each database is then pushed to its own Github repository, hosted by the metapsy-project account.
- Database repositories are then officially released using Zenodo. At this step, a unique version number and digital object identifier (DOI) is created.
- The database documentation hosted on this website is automatically updated with each new release. The documentation entry contains metadata and additional information for each database.
- The
metapsyDataR package allows to access all Metapsy databases directly in an R environment. JSON-encoded exports can be retrieved using the Metapsy API. - The
metapsyToolsR package provides helper functions that allow to conveniently run state-of-the-art meta-analyses using Metapsy data, or data prepared in the same format. - The released databases can also be analyzed using a user-friendly meta-analysis tool. The application provides a graphical user interface that allows running meta-analyses of the data online.