R Packages
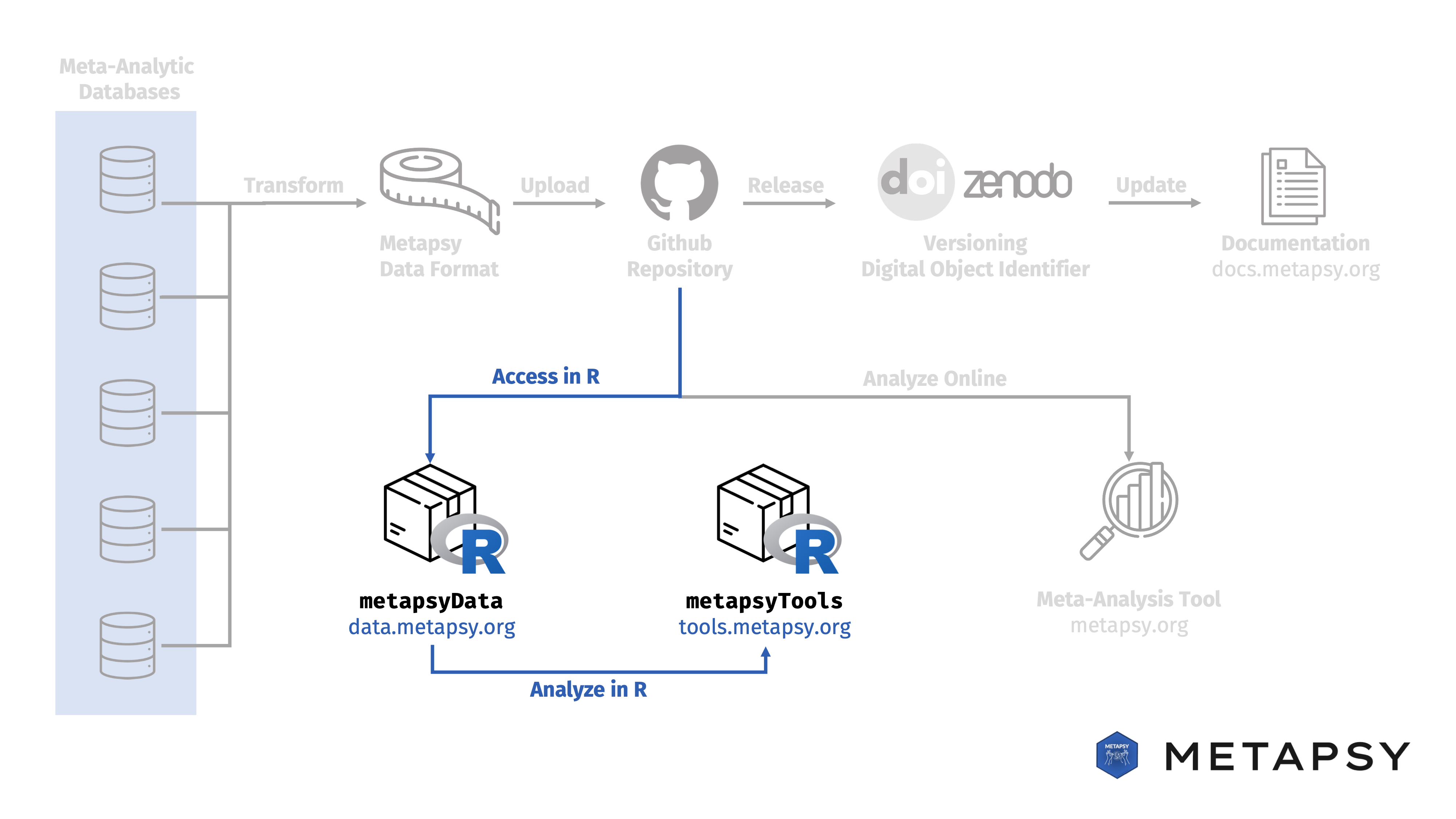
The Metapsy infrastructure includes two R packages. The first package, metapsyData, allows researchers to directly access all Metapsy databases from an R environment. The second package, metapsyTools, provides state-of-the-art meta-analysis functions that can be applied to databases without any prior preprocessing steps. Collectively, these packages create an integrated meta-analysis workflow. Using the packages:
- 📥 the latest update or older versions of a database can be downloaded;
- 👉 databases can be filtered to address specific research questions;
- 📊 data can be synthesized using a variety of meta-analytic models;
- 📈 meta-regression, small-study-effect and subgroup analysis models can be applied;
- 📋 summary tables of the results can be generated, which can easily be copied to e.g. MS Word.
metapsyData
The metapsyData package serves as an API-like interface that allows accessing the Metapsy databases from R. Using the getData() function, available databases can be downloaded based on their shorthand. The function returns an R6 metapsyDatabase object, which contains extensive metadata and helper functions along with the actual dataset.
Using the listData() function in metapsyData, it is possible to list all datasets that are currently available, as well as their shorthands. Databases and shorthands are also presented on the Metapsy website.
The metapsyData package has its own documentation page, which you can find here.
metapsyTools
The metapsyTools package facilitates the calculation of effect sizes (i.e. Hedges’ $g$ or risk ratios) and meta-analyses for data included in Metapsy databases (or databases adhering to the same format).
The package consists of two modules:
- A module to check if data follows the Metapsy data standard, and to calculate effect sizes for all possible study comparisons (preparation module);
- A module to select relevant comparisons for a specific meta-analysis, calculate the results (including subgroup analyses, meta-regression, small-study-effect/publication bias analyses), and generate tables (analysis module).
The idea is to use the two modules in different contexts. For example, the preparation module can be used every time the database is updated to gather all information, calculate effect sizes, and bring the data into a format suitable for further analyses.
This final dataset then builds the basis for the analysis module. Researchers simply have to filter out the comparisons that are relevant for their investigation, and can then use functions of the package to run a full meta-analysis.
Databases downloaded into the R environment using metapsyData can be analyzed “out of the box” using the analysis module in metapsyTools.
Like metapsyData, the metapsyTools package has its own documentation page. You can find it here.
Data Validator
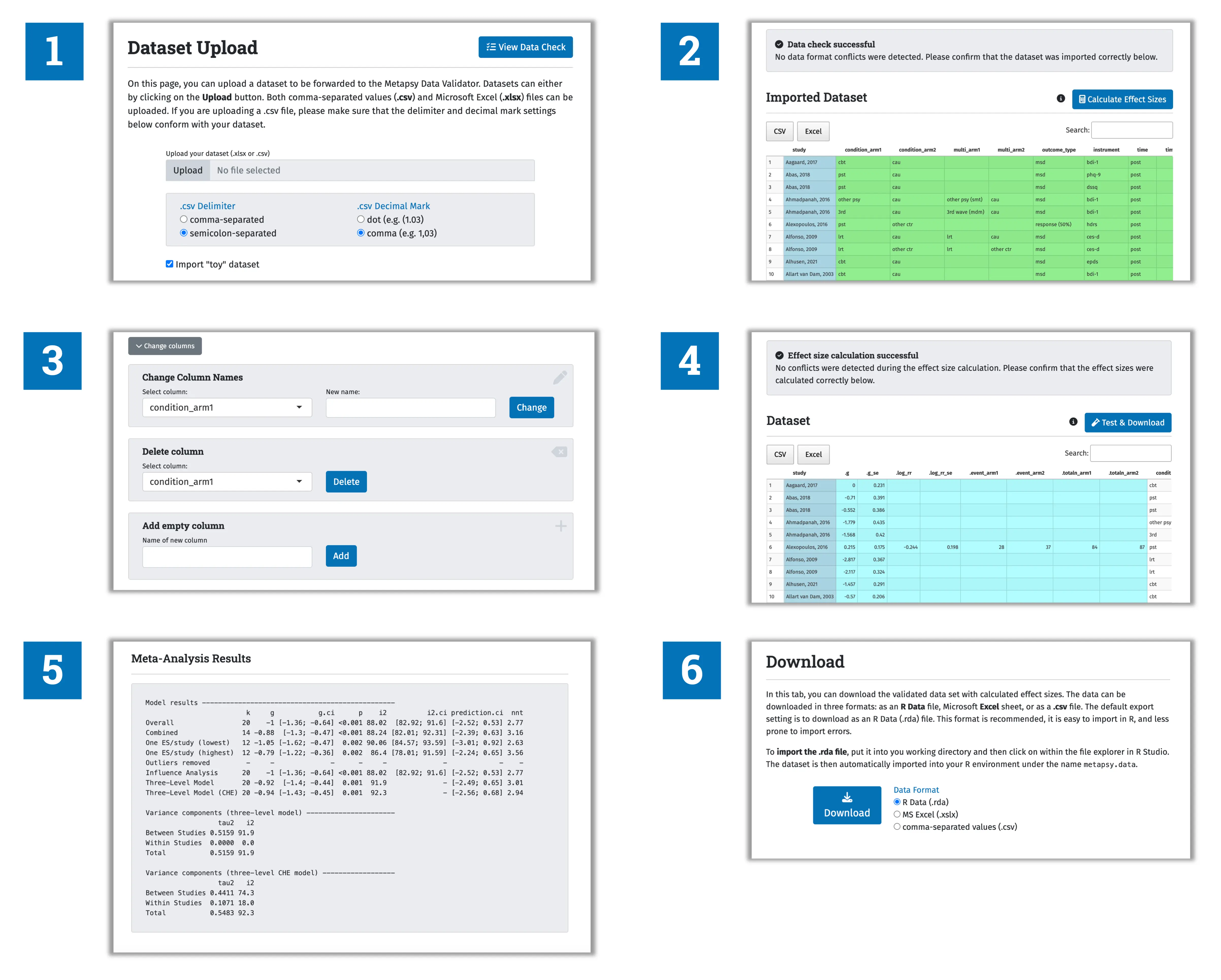
The preparation module of the metapsyTools package has also been implemented in a digital web application, the Metapsy data validator. This tools allows to upload an existing dataset whose compatibility with the Metapsy data standard should be checked (see 1 in the figure below).
The application automatically runs a test to identify potential formatting issues (2), which can be resolved using the graphical user interface (3). The tools then calculates all effect sizes for the dataset and saves them in the correct format (4).
Lastly, users can check if a meta-analysis can be run with metapsyTools using the formatted data (5), and it is possible to download the final dataset (6).